Supporting research on Covid-19
How neutron science supports research on viruses such as Covid-19
A lot of action in the virus’s life cycle happens at the surfaces. Like a space ship docks onto a space station the virus has to hook onto the cell it intends to infect and create an interlock for dumping its genetic material. In the specific case of SARS-CoV-2, which looks very much like a sea mine, the proteins of the spikes dock onto the epithelial cells in the patient’s airways by binding a special enzyme (ACE2) expressed – and therefore present - on those cells’ surface.
A considerable part of research on SARS-CoV-2 will be targeting these docking processes. Neutron reflectometry, which is the neutron technique sensitive to surfaces, can tell you how it happens - like in the case of the hepatitis C virus research performed in 2017. ILL possesses a top-of-the-range reflectometry suite and outstanding expertise in applying this suite to the study of membrane systems.
Press release
Press cuttings
Scientific publications
- Coupling neutron reflectivity with cell-free protein synthesis to probe membrane protein structure in supported bilayers | Scientific Reports (2017)
FIGARO experiment to study the interaction of p7 Viroporin, a membrane protein involved in the assembly and release of hepatitis C virus. - Cholesterol modulates the fusogenic activity of a membranotropic domain of the FIV glycoprotein gp36 | Soft Matter (2013)
- Fusion of raft-like lipid bilayers operated by a membranotropic domain of the HSV-type I glycoprotein gH occurs through a cholesterol-dependent mechanism | Soft Matter (2015)
- Destabilization of Lipid Membranes by a Peptide Derived from Glycoprotein gp36 of Feline Immunodeficiency Virus: A Combined Molecular Dynamics/Experimental Study | J. Phys. Chem. B (2012)
Neutron crystallographyis one of the indispensable modern analytical tools for obtaining insight into the life cycle of viruses.
In simple terms, crystallography provides us with three-dimensional pictures of the various biochemical engines that the virus relies upon for reproduction. If these engines can be blocked through appropriate medication e.g. by introducing a molecule into the site where the biochemical action takes place, then the viral disease can be cured.
Many of these engines manipulate hydrogen nuclei. This is where the neutrons come in. They are ideal for detecting hydrogen nuclei in the structure. The ILL is the leading neutron facility in the domain of biological macromolecular neutron crystallography. Recent studies on HIV-1 protease (a biochemical engine that - like a pair of scissors - cuts long polymer chains and is essential for the life-cycle of the HIV virus) performed at the ILL’s instrument LADI perfectly illustrate the case. The neutron data allows to better design the drug that blocks the scissors in their active site.
SARS-CoV-2 offers many potential targets for such neutron studies. Given the relevance of this research, ILL has decided -in the framework of the Endurance upgrade programme- to enhance capacity by adding another instrument (DALI) (pdf - 2.36 Mi) to its instrument suite. DALI was just in the process of being assembled when the lock-down hit the ILL activities. The installation will be completed after the lock-down and the instrument will be commissioned during the next cycle.
Press releases
Press cuttings
- Neutrons key to discovering new HIV drugs? An interview with Dr Matthew Blakeley | News-medical.net
- Neutrons probe structure of enzyme critical to development of next-generation HIV drugs | Phys.org
- Fighting HIV with neutrons | Science in School
- Neutron study aims to improve HIV drugs | Physics World
- In the Spotlight | European Biopharmaceutical Review
- Neutron Studies Reveal New Areas for Improvement in Drug Design of HIV Inhibitors | PharmTech.com
- Neutron crystallography aids in drug design | ScienceDaily.com
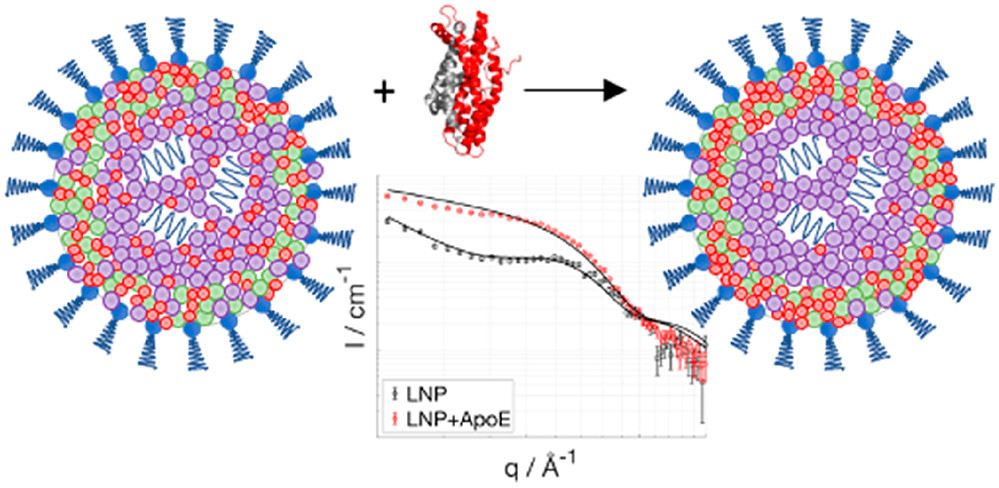
When it comes to studying the function of larger biological complexes such as assembled viruses, small angle neutron scattering becomes an important analytical tool. This is due to the fact that: the larger the objects under investigation, the smaller gets the angle of deviation of the neutrons during scattering from those objects. Neutrons offer the enormous advantage that individual subunits can be marked by advanced deuteration allowing to distinguish specific regions (RNA, proteins and lipids) in the complexes.
While NMR and cryo-electron microscopy provide the detailed atomic-resolution structure of small biological assemblies, neutron scattering allows researchers to pan back to see the larger picture of full molecular complexes at lower resolution. Neutron scattering is also uniquely suited to determining the structure of functional membrane proteins in physiological conditions. Neutron scattering will therefore make it possible to map out the structure of the complex formed by the SARS-CoV-2 spike protein—the protein surrounding the virus—and its human receptor.
With its excellent suite of small-angle instruments and the outstanding expertise in applying them to biological systems, ILL is perfectly positioned for performing such studies on SARS-CoV-2.
Press cuttings
An even closer look at the COVID-19 infection process (pdf - 258 Ki) | GIANT Review Spring 2022
Scientific publications
- Apolipoprotein E Binding Drives Structural and Compositional Rearrangement of mRNA-Containing Lipid Nanoparticles | ACS Nano (2021)
- RNA Back and Forth: Looking through Ribozyme and Viroid Motifs | Viruses (2019 )
- Ensemble Structure of the Highly Flexible Complex Formed between Vesicular Stomatitis Virus Unassembled Nucleoprotein and its Phosphoprotein Chaperone | Journal of Molecular Biology (2016)
- Large-Scale Conformational Dynamics Control H5N1 Influenza Polymerase PB2 Binding to Importin α | J. Am. Chem. Soc. (2015 )
- Intrinsic disorder in measles virus nucleocapsids | PNAS (2011)
- Structure and dynamics of the nucleocapsid-binding domain of the Sendai virus phosphoprotein in solution | Virology (2004)
- Combined results from solution studies on intact influenza virus M1 protein and from a new crystal form of its N-terminal domain show that M1 is an elongated monomer | Virology (2001)
- On the domain structure and the polymerization state of the Sendai virus P protein | Virology (2000)
Finally, we should not forget that viruses in their physiological environments are highly dynamic systems. Knowing how they move, deform and cluster is essential to the optimisation of diagnostic and therapeutic processes. Neutron spectroscopy, which is ideally suited to follow the motion of matter from the small chemical group to large macromolecular assemblies, is the tool of choice to provide this information.
Scientific publications
- Proton transfer and drug binding details revealed in neutron diffraction studies of wild-type and drug resistant HIV-1 protease | Methods in Enzymology (2020)
- Protein kinase A in the neutron beam: Insights for catalysis from directly observing protons | Methods in Enzymology (2020)
- Zooming in on protons: Neutron structure of protein kinase A trapped in a product complex | Science Advances (2019)
- A molecular mechanism for transthyretin amyloidogenesis | Nature Communications (2019)
- The helix-to-sheet transition of an HIV-1 fusion peptide derivative changes the mechanical properties of lipid bilayer membranes | Biochimica et Biophysica Acta (BBA) - Biomembranes (2018)
- Elucidation of hydrogen bonding patterns in ligand-free, lactose- and glycerol-bound galectin-3C by neutron crystallography to guide drug design | Journal of Medicinal Chemistry (2018)
- “To be or not to be” protonated: Atomic details of human carbonic anhydrase-clinical drug complexes by neutron crystallography and simulation | Structure (2018)
- The GAIIG domain from Amyloid-β and the HIV-1 gp120 V3 loop as a Source of Inspiration for Novel Amyloid Scaffolds and Potential Therapeutics | FEBS Letters (2018)
- Room temperature neutron crystallography of drug resistant HIV-1 protease uncovers limitations of X-ray structural analysis at 100K | Journal of Medicinal Chemistry (2017)
- Direct visualization of critical hydrogen atoms in a pyridoxal 5′-phosphate enzyme | Nature Communications (2017)
- Long-range electrostatics-induced two-proton transfer captured by neutron crystallography in an enzyme catalytic site | Angewandte Chemie International Edition (2016)
- Human CD4 Metastability Is a Function of the Allosteric Disulfide Bond in Domain 2 | Biochemistry (2016)
- Self-Assembly of an Aspartate-Rich Sequence from the Adenovirus Fibre Shaft: Insights from Molecular Dynamics Simulations and Experiments | J. Phys. Chem. B (2014)
- Neutron diffraction reveals hydrogen bonds critical for cGMP-selective activation: Insights for cGMP-dependent protein kinase agonist design | Biochemistry (2014)
- Binding site asymmetry in human transthyretin: insights from a joint neutron and X-ray crystallographic analysis using perdeuterated protein | IUCrJ (2014)
- Joint X-ray/neutron crystallographic study of HIV-1 protease with clinical inhibitor amprenavir – insights for drug design | Journal of Medicinal Chemistry (2013)