Room to move: how proteins behave in crowded environments
21 Apr 2026Neutron scattering shows how local density shapes motion within self-assembled protein structures
-
Health

Inside living cells, proteins are constantly moving, interacting and organising into complex structures. In many cases, they gather into dense assemblies where space is limited and motion becomes constrained. Understanding how proteins behave in such crowded environments is essential for explaining key biological processes, from cellular organisation to the formation of protein aggregates linked to disease.
Using neutron scattering at the Institut Laue - Langevin, combined with computer simulations, researchers studied how a particular type of protein, behaves inside dense assemblies formed by the proteins themselves. They found that the motion of these proteins is not uniform: proteins move more slowly in the dense centre of the assemblies and more freely near the edges. This shows that how proteins organise themselves directly affects how they move, an effect that is important for understanding how biological systems function.

A self-created crowded environment
Proteins are essential molecules in living systems. They move, interact and organise themselves to carry out a wide range of functions, from helping cells communicate to forming structures inside the cell. In many cases, proteins do not remain isolated, but instead spontaneously group together, forming assemblies made of many individual molecules. This process, known as self-assembly, is driven by interactions between the proteins themselves.
These assemblies create environments that are very different from simple liquids. Instead of moving freely, proteins are surrounded by many neighbours, and space can become limited. Such crowded conditions are very common inside cells and can strongly influence how molecules move and interact. Understanding how proteins behave in these environments is therefore essential for describing how biological systems function.
In this study, researchers investigated how a flexible protein called β-casein behaves inside such assemblies using neutron scattering at the Institut Laue-Langevin (ILL), combined with computer simulations. Because these assemblies are formed by the proteins themselves, they create a “self-crowded” environment in which local conditions vary from one region to another. β-casein is an example of an intrinsically disordered protein, meaning it does not have a stable structure and remains highly flexible.
Neutrons reveal complex motion at the nanoscale
To observe these effects directly, the team used quasielastic neutron scattering (QENS) on the IN16B spectrometer at the ILL. This technique measures very small changes in neutron energy as they interact with moving atoms in the sample, allowing researchers to track how molecules move on extremely short timescales (from picoseconds to nanoseconds, or trillionths to billionths of a second) and over very small distances (nanometres, a million times smaller than a millimetre).
Neutrons are particularly well suited for studying proteins because they are highly sensitive to hydrogen atoms, which are abundant in these molecules. This makes it possible to probe the motion of individual protein chains even inside dense assemblies, where other techniques struggle to access such information.
In a simple picture known as Fickian diffusion, molecules move randomly but in a predictable overall way:: although they tend to spread from regions where they are more concentrated to regions where they are less concentrated, their motion can still be described by a single, uniform rule across the whole system.
In the assemblies of the disordered proteins studied here, the neutron measurements reveal a different picture. Instead of a single, uniform type of motion, the data show a deviation from Fickian diffusion and instead follow a behaviour known as Singwi-Sjölander diffusion, indicating that the protein motion depends on their immediate surrounding within the same assembly. This behaviour is directly reflected in the neutron signal, which cannot be described by a single, simple diffusion process.
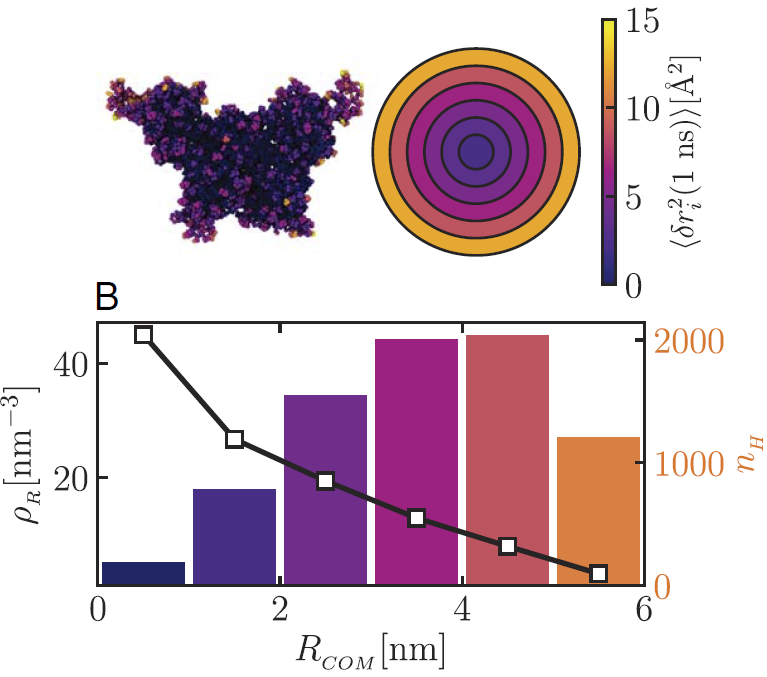
Motion inside a protein assembly is not uniform. (Top left) The assembly is coloured according to how much the proteins move: motion is slower in the dense centre (dark colours) and faster toward the edges (yellow). (Top right) A simplified view showing how this motion changes gradually from the centre to the outside. (Bottom) The distribution of protein material as a function of distance from the centre of the assembly (RCOM). The black line shows how densely packed the proteins are (ρR), with the centre being much more crowded than the outer regions. The coloured bars show the number of hydrogen atoms (nH) in each region, indicating where most of the protein material is located. Together, this shows that proteins are more tightly packed in the centre, which limits their motion, while those near the edges have more space and can move more freely. Credit: Proc. Natl. Acad. Sci. U.S.A. (2026).
Computer simulations help explain the origin of this behaviour by linking the observed motion to the internal structure of the assemblies. They show that the assemblies are not uniform: the centre forms a dense, tightly packed region, while the outer parts are less crowded. This structural variation leads directly to differences in mobility, with slower motion in the dense core and more freedom near the edges. Importantly, this change is gradual: motion evolves continuously from the centre toward the outer regions, rather than switching abruptly between two distinct behaviours.
This behaviour, known as non-Fickian diffusion, shows that the internal organisation of the assembly directly shapes how proteins move. Because motion varies across the structure, it cannot be described by a single diffusion law, illustrating the limits of simple models for describing molecular motion in crowded biological environments.
From structure to motion
These findings demonstrate how the organisation of proteins at the nanoscale can directly influence how they move and interact. Such effects are likely to be important in living cells, where molecules rarely operate in isolation but instead function in crowded and structured environments.
Beyond this specific system, the results highlight the limits of simple diffusion models for describing molecular motion in complex media. When local structure varies, as it does in many biological and soft-matter systems, motion can no longer be captured by a single, uniform description.
By revealing these effects experimentally, neutron scattering provides a powerful way to connect molecular organisation with dynamics, helping to build a more realistic picture of how biological systems function at the microscopic level.
Reference:
L.M. Miñarro, S. Chakraborty, C. Beck, A.C. Grundel, I. Mosca, F. Roosen-Runge, T.I. Morozova, J. Barrat, F. Schreiber, & T. Seydel, Non-Fickian diffusion within assemblies of the intrinsically disordered protein β-casein, Proc. Natl. Acad. Sci. U.S.A. 123 (11) e2532636123, (2026). DOI: 10.1073/pnas.2532636123
ILL instruments: IN16B
ILL Contact Person: Tilo Seydel
Institutions involved in the research: Université Grenoble-Alpes, University of Tübingen, Heidelberg University, Lund University and École Normale Supérieure de Lyon